AI Improves Quality Control of Cell- and Gene Therapy Products
A Glance Into the Future of QC for ATMPs—In the 1980s, advancements in genetic engineering have revolutionized the pharmaceutical industry and created a new category of medicines: biopharmaceuticals. This development has led to numerous ground-breaking treatments for autoimmune diseases and cancer, and has laid the foundation for RNA vaccines currently developed against Covid-19.
In the last decade, technological advancements have created yet another new category of medicine that has the potential to impact the pharmaceutical industry in a similar way: cell-based medicinal products (CBMPs). CBMPs belong to the large group of advanced therapy medicinal products (ATMPs) and are based on living cells from the patient itself or a healthy donor (read a commentary about the term ATMP). In most cases, these cells are genetically modified (by e.g., viral vectors) to exert their therapeutic effect.
Quality Control is Critical for CBMPs
A prominent example of a CBMP is used for treating cancer: Chimeric antigen receptor T-cells, also called CAR T-cells. Generally, T-cells control and direct immune responses and are a vital part of our immune system. During CAR T-cell immunotherapy, naïve (=immature) T-cells are harvested from the patient and transferred to a lab for genetic modification. The modified genes add tumor-specific receptors to the surface of the T-cells and, when administered back into the patient, the CAR T-cells will induce an immune response against the tumor.
As for all medicinal products, such personalized medicine needs to fulfill certain quality criteria. Their universal implementation for CBMPs is one of the key areas, which is being extensively reviewed to ensure patients’ safety. In the literature, the difficulty in the application of pharmaceutical quality control to CBMPs is frequently mentioned. The complexity and stability issues of cells makes analysis of product comparability, stability and compatibility extremely challenging. Several draft guidelines (i.e., EMA/CAT/852602/2018) have been recently published for public consultation aiming to consolidate knowledge in the field for establishing improved strategies in the development of CBMPs.
“Cellular impurities, such as cell aggregates and adducts, can be easily classified by using a our neural network approach.”
Adam Grabarek, Study Author
Quality control of CBMPs includes testing for the absence of particulate impurities. For non-cell-based medicines, one can simply remove undesired particles through low pore-size filtration. But since living cells in CBMPs can be as large as 30 µm (T-cells are around 10 µm in size), filtration is not a valid option. Of concern are micrometer-sized particles such as Dynabeads (4.5 µm) that are added to the CBMP during T-cell activation. Due to the risk of a certain toxicity, their absence (after removal via magnetism) must be confirmed analytically.
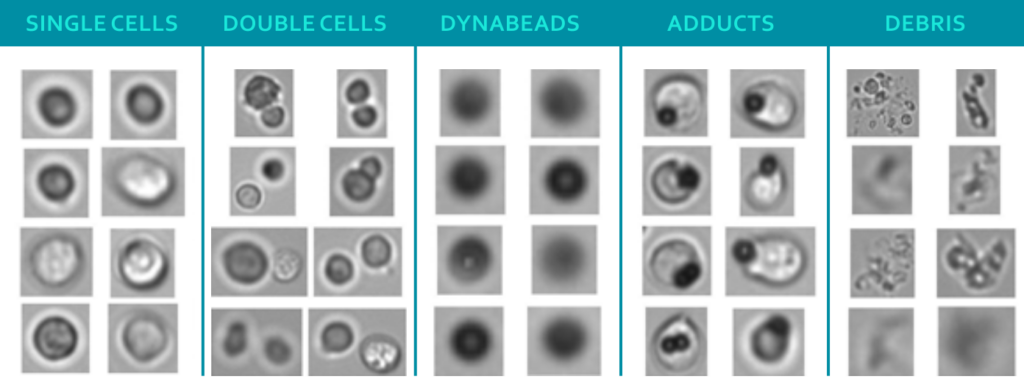
The figure shows images of particles in a CAR T-cell sample taken by flow imaging microscopy (FIM), a technique for sizing, quantifying and visualizing particles. The single cells are the desired compound of the CBMP, while Dynabeads and adducts (agglomeration of Dynabeads and cells) are undesired. Current quality control protocols utilize manual microscopy with low throughput and a small analyzed sample volume. On top of that, the time between production and administration of the drug product, during which quality controls takes place, is rather short. This led to the desire for new and improved analytics. That is why researchers from Coriolis Pharma, in collaboration with the Leiden Academic Centre for Drug Research (LACDR, The Netherlands) and the Leiden University Medical Center (LUMC, The Netherlands) explored ways of improving speed, accuracy and throughput of analyzing particles in CBMPs.
The research groups around lead scientist Adam Grabarek used Jurkat cells and Dynabeads to obtain representative bright field images of all particle types shown in the figure. To classify the images into single cells, Dynabeads and adducts, the researchers used a manual, as well as a machine learning approach based on convolutional neural network (CNN) approach. For the manual approach, the team applied statistical analysis of morphological parameters, such as size, intensity, aspect ratio and convexity for classification. For the CNN approach, the scientists applied transfer learning of a pre-trained VGG-19 network, which was refined with a dataset of labelled particle images.
Artificial Intelligence Outperforms Manual Approach
Particle characterization classification cell-based medicinal products CBMPThe researchers could show that the manual approach correctly identified ca. 90% of single beads and ca. 88% of cells in samples consisting of a single population (only beads or only cells). However, the ability to correctly identify beads in samples of a mixture of cells, beads and debris decreased dramatically. By contrast, the CNN approach was able to correctly identify >98% of beads and cells in samples with single particle populations and the accuracy remained similar for samples consisting of particles from all classes presented in the figure above. The team recently published all their results in the Cytotherapy Journal.
“In our study, we developed a reliable method based on FIM coupled with CNN for the detection, characterization and quantification of relevant particulate impurities, specifically Dynabeads.” said Adam Grabarek. “We showed that small amounts of Dynabeads can be detected in cell suspensions and a high precision in counting is achieved […]. Moreover, cells and cellular impurities, such as cell aggregates and adducts, can be easily classified by using CNN.”
Further studies are currently being conducted to enhance the capabilities of flow imaging in combination with machine learning to other cell lines and particle types. The research group is also hoping for other teams to evaluate their method. This could increase the confidence in and understanding of the analytical approach and may support its use in product development as well as support manufacturing in-process controls. “Currently, we are not aware of other methods with similar performance,” explained Adam Grabarek, ”and we believe that CBMP development can benefit from our method in its current state.”
